The researchers who first reported the Omicron variant have published their findings about the discovery of Omicron a few days ago. I will explain how they discovered Omicron by summarizing some of the findings from their publication1.
A recap of Omicron
Almost two months ago, I published a blog post about the first reports of Omicron from South Africa. Before Omicron emerged, South Africa had already experienced three waves of SARS-Cov-2, each driven by different lineages over time. As of today, Omicron has been detected in over 120 countries in every continent. Since then, much has been written about Omicron, ranging from how well vaccines work against Omicron (eg. booster shots can provide potent neutralization against omicron2) to Omicron inducing less severe COVID-19 disease than Delta3. This blog post won’t go into more detail about these findings, but will instead focus on painting a picture of how Omicron was first detected and characterized.
How was Omicron identified?
In mid-November 2021, a sudden rise in COVID-19 cases was detected in Gauteng Province. Gauteng Province is home to the largest city in South Africa, Johannesburg, and accounts for 25% of the country’s total population despite being the smallest province in the country (Figure 1). The country had just recently ended the third wave driven by Delta a month before, so could the new November wave be driven by Delta again? We have polymerase chain reaction (PCR) tests that can detect SARS-Cov-2 by amplfiyng its genetic material. However, just targeting the virus alone cannot shed light to the lineage(s) (ie. Alpha, Beta, Delta, etc.) that’s infecting a person with COVID-19.

Thankfully, PCR tests don’t have to be limited to identifying just a single target. One way the authors could start knowing which lineage was infecting a person was to perform a series of polymerase chain reaction (PCR) assays at once, using a TaqPath COVID-19 test kit (ThermoFisher Scientific)4. The primary strength of this test kit lies in its ability to identify the presence of SARS-Cov-2 in a person by targeting three regions of the SARS-Cov-2 genome (Figure 2). Two of these regions are essential for SARS-Cov-2’s functioning: open reading frame (ORF) ORF1ab and the nucleocapsid. The function of ORF1ab is unknown, but the nucleocapsid packages the RNA-based genetic material into a package within the virus. The last of these regions is the spike protein (S), so critical for invading the human lungs and causing COVID-19 in the first place. Each of these regions can be targeted at the same using the TaqPath COVID-19 test through multiplex PCR tests that measure the amount of SARS-Cov-2 genetic material in our airways.
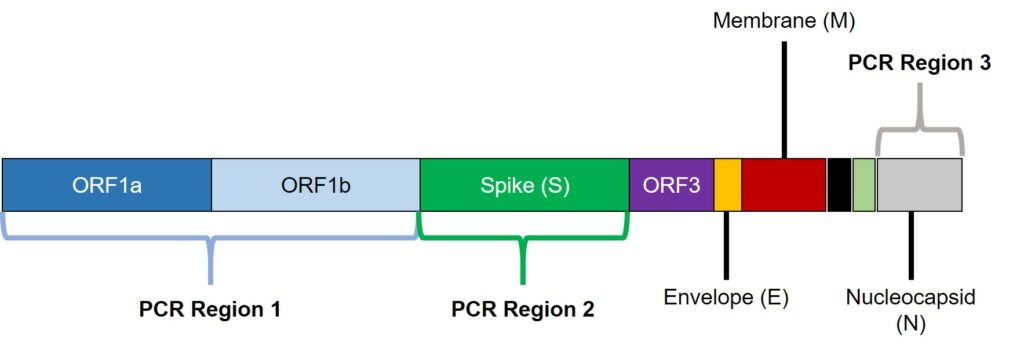
Although the ORF1ab and nucleocapsid PCR targets are essential for detecting SARS-Cov-2, the spike protein PCR assay was key to identifying the Omicron variant. The Taqpath PCR assay actually relies on not being able to detect the spike protein to identify variants with spike protein mutations. This assay was first developed to identify the Alpha variant which had a deletion to the spike protein at amino acid positions 69 and 70. If you recall my previous blog post, the Omicron variant has 32 mutations to the spike protein alone. Among those mutations is a deletion at amino acid positions 69 and 70, which was also observed in the Alpha variant and not the Delta variant. With the fact that the Delta variant beat out the Alpha variant in South Africa, the authors considered the possibility of a new variant being identified through their PCR assays.
With that in mind, the SARS-Cov-2 genomes with the characteristic deletion mutations in the spike protein were then sent for whole genome sequencing to see if the strain was a new variant. As recent history would have it, the Omicron variant was then discovered, with its 32 mutations to the spike protein. Interestingly, a series of genomes sequenced from samples obtained in Botswana also contained the same sets of unusual mutations. Each of those samples were linked to a group of visitors who were on a diplomatic mission. From then on, all the genomes were considered to the part of the B.1.1529 lineage that we now know as Omicron.
So where did Omicron come from?
At the time of the study’s publication, 686 Omicron genomes were sequenced, 248 from South Africa and 438 from the rest of the world. These genomes were compared against the 12609 representative SARS-Cov-2 genomes obtained worldwide. If you observe Figure 2 and Extended Data Fig. 4 in the paper, you will see a series of phylogenetic trees. A phylogenetic tree is like a family tree, but for animals and microbes. It helps scientsts like me trace how microbes like SARS-Cov-2 mutated over time and where they reproduced from, given that we have the full genomes and the time the samples were collected. Each of those genomes are represented by a terminal node attached to a branch. If a group of specimens cluster together within the tree, it means we can trace those branched specimens back to a single recent common ancestor at an internal node. Those ancestors can then be traced back further along the branch and so forth all the way back to the root. This can be done for every organism, regardless of where in the tree they belong.
Now with that in mind, if you see the phylogenetic trees again, you’ll see two things. First, you’ll see that the genomes belonging to Omicron are clustered together away from the other variants within the tree. That makes it less likely that Omicron came from a specific lineage. More interestingly, there seems to be three distinct sublineages of Omicron, called BA.1, BA.2, and BA.3. In addition, BA.2 and BA.3 seem to have evolved independently of the BA.1 sublineage. That would explain the BA.2 and BA.3 genomes are in a completely different branch from the BA.1 sublineage. The Omicron variant predominating around the world seems to be the BA.1 sublineage, but I can’t help but wonder where the BA.2 and BA.3 sublineages came from or what’s become of it. Perhaps with so many Omicron cases maybe we’ll get a better picture of where BA.2 and BA.3 Omicron comes into play.
Nevertheless, I think it’s gonna be a very hard challenge to know precisely where Omicron came from. After all, the scientists only had 686 Omicron genomes at the time of the study in a large sea of Omicron cases in South Africa when it first emerged. Further complicating matters is the fact that Dalhousie researchers also identified Omicron after rechecking their wastewater samples from mid-November with PCR, before it was first identified in South Africa5. The results certainly make it more difficult to know where Omicron really came from since it’s likely that Omicron was already spreading even before it was first identified in South Africa. In short, I don’t think we can answer this question right now, even if hypotheses are being generated regarding their origins6.
So what next?
The rest of the paper spends time talking about the nature of the mutations in the Omicron variant and the role that mutational adaptation plays in the emergence of Omicron. It’s clear that Omicron;’s mutations are being selected for according to the scientists. Much of the selection for these mutations is occurring at the spike protein, which has been the primary target for vaccines to fight off SARS-Cov-2 infection. I imagine it’s a big reason why COVID-19 vaccines are less effective at two doses and why boosters are more effective against Omicron. But how sustainable is that? Much of the world doesn’t even have two doses of the vaccine yet. Even if a South African person gained immunity from natural infection, an exponential increase in Omicron cases was still observed among South Africans. That also suggests that Omicron can partially evade immune responses against SARS-Cov-2. All these variables play a role in Omicron’s exponential rise in cases for much of the world, including Canada and the USA.
That all being said, let’s hope that Omicron cases will go down soon as more people get booster shots or infected. That is what we’re seeing in South Africa so far after they’ve had an Omicron wave for 1-2 months.
References
- Viana, R., Moyo, S., Amoako, D.G., Tegally, H., Scheepers, C., Althaus, C.L., Anyaneji, U.J., Bester, P.A., Boni, M.F., Chand, M., et al. (2022). Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature.
- Garcia-Beltran, W.F., St Denis, K.J., Hoelzemer, A., Lam, E.C., Nitido, A.D., Sheehan, M.L., Berrios, C., Ofoman, O., Chang, C.C., Hauser, B.M., et al. (2022). mRNA-based COVID-19 vaccine boosters induce neutralizing immunity against SARS-CoV-2 Omicron variant. Cell S0092-8674(21)01496-3.
- Sohn, Emily. January 11th, 2022. National Geographic. https://www.nationalgeographic.com/science/article/is-omicron-really-less-severe-heres-what-the-science-says
- Clinical Conversion Staff. January 14th, 2021. ThermoFisher Scientific. https://www.thermofisher.com/blog/clinical-conversations/the-s-gene-advantage-taqpath-covid-19-tests-may-help-early-identification-of-b-1-1-7/
- Mundie, Jessica. January 10th, 2022. National Post. https://nationalpost.com/news/canada/omicron-was-in-nova-scotia-wastewater-before-it-was-identified-in-south-africa
- Haseltine, William A.; December 2nd 2021. Forbes. https://www.forbes.com/sites/williamhaseltine/2021/12/02/omicron-origins/#:~:text=The%20three%20most%20likely%20origin,with%20the%20mutagenic%20drug%20molnupiravir.
